smart toilet paper
a simple, disposable device to track your gut microbiome
Worldwide, there are about 3-5 billion cases of diarrhoeal diseases, many of which may be classified as gastroenteritis.[1] If gastroenteritis are not treated early, it may lead to further complications such as Crohn’s Disease, Ulcerative Colitis, Irritable Bowel Syndrome, Reactive Arthritis, and other Arthritis.[2] The risk for developing these diseases can be assessed through a microbiome analysis. The instruments used to measure the microbiome exist, however they are complex and expensive and therefore inaccessible for a large section of the global population.
Can we produce a simple and affordable device that can effectively track the microbiome in the human gut?
FECAL TO DATA
What do we need to do to know if we have a certain type of bug in your gut? Fecal samples are collected. The microbial DNA within the samples are extracted, amplified, and visualized.
THE SCIENCE
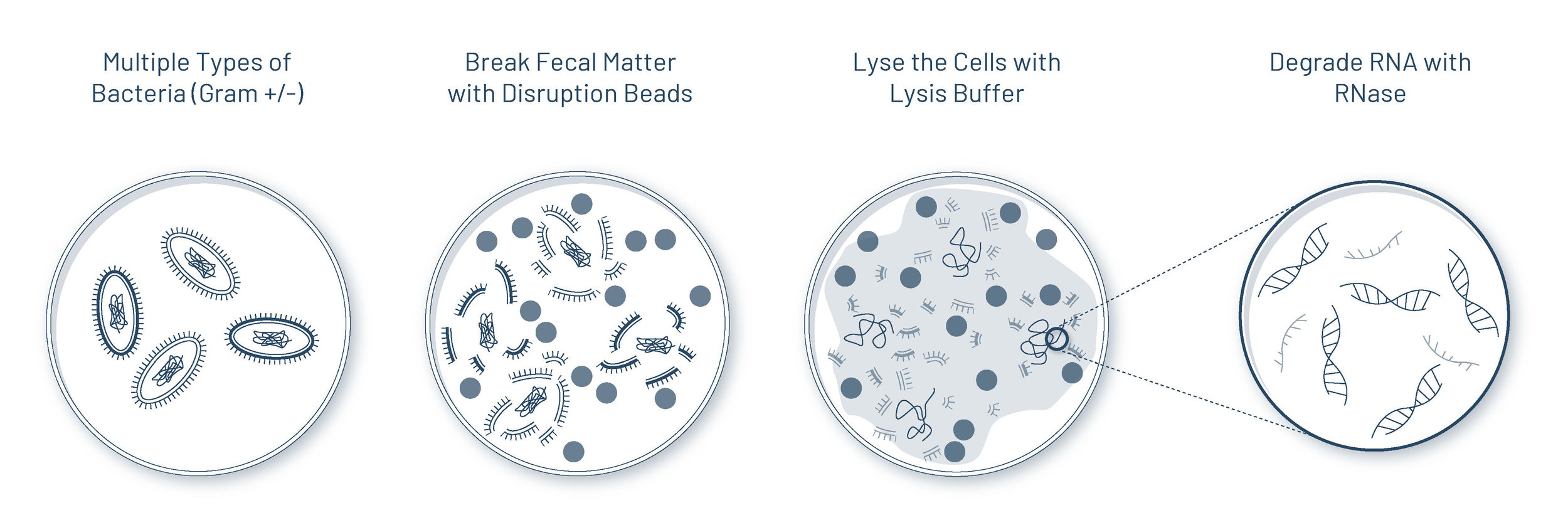
Since there are many types of bacteria, including both gram-negative bacteria and gram-positive bacteria (those with much thicker walls), disruption beads are used to mechanically break them open. The lysis buffer, a detergent-based buffer solution, further breaks open the cells and isolates the different cellular components. RNase is then introduced to degrade RNA to better isolate for DNA.
DNA Extraction through the spin column
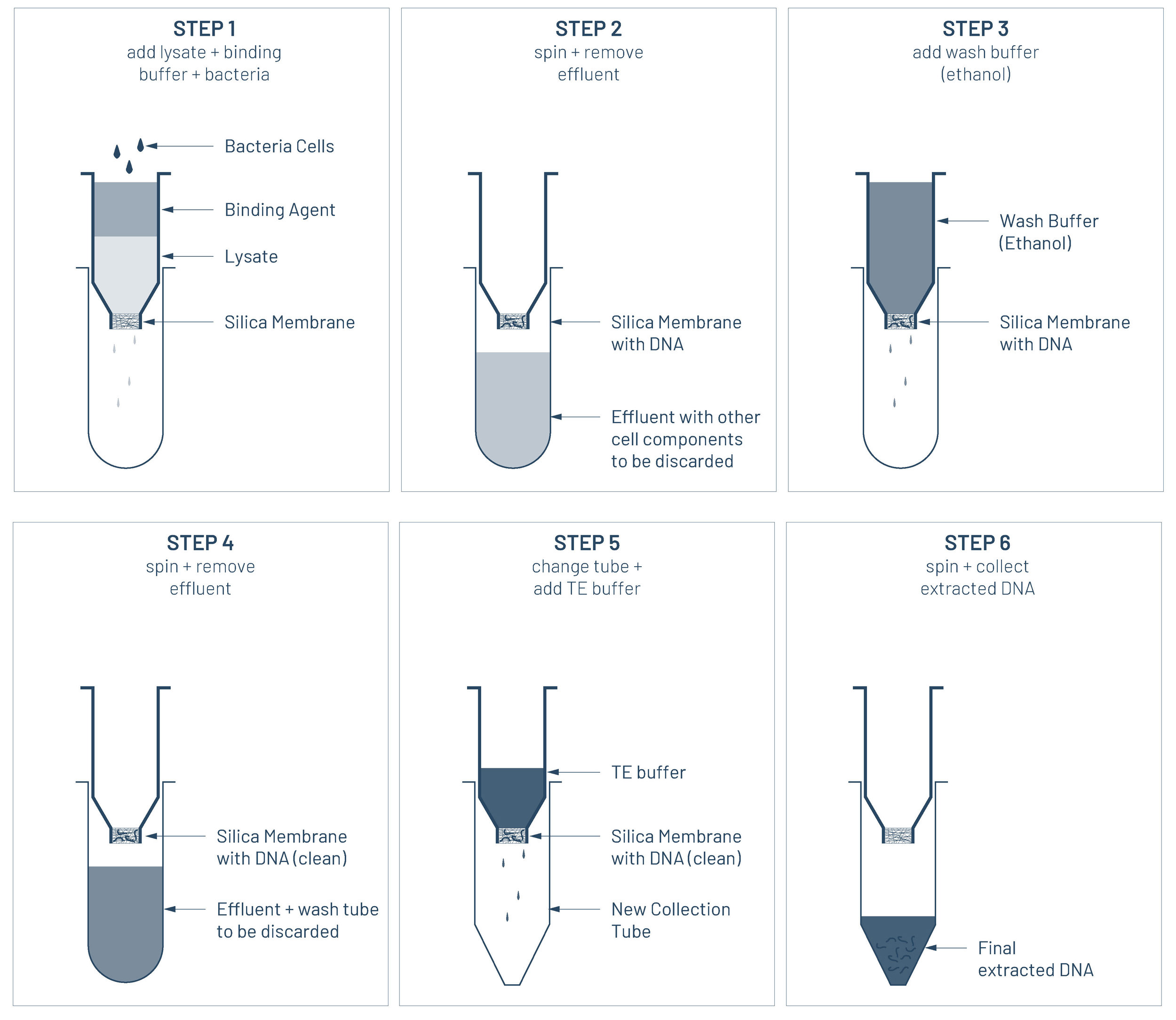
The method used is known as the spin-column method. A spin column is a tube with a small opening at the bottom that is filled with silica membrane. It takes advantage that DNA (which is negatively charged on its surface), tend to bind to silica (glass) membranes or beads (which have positively charged surfaces), when they’re in the presence of certain salts and at a low pH level. TE buffer is added to the spin column to then remove the cleaned DNA from the silica and extracted in a tube.
Prototypes + PROCESS
The realization of the project was multifaceted, touching on geometry, physics, bio-chemistry, fabrication, material exploration, and user experience.
More detailed information on the project upon request.
REFERENCES
[1] https://my.clevelandclinic.org/health/diseases/12418-gastroenteritis
[2] https://www.badgut.org/information-centre/a-z-digestive-topics/gastroenteritis/